Genomics Capabilities

Single cell analysis is essential to detecting and understanding cancer heterogeneity.
The Epic Sciences' platform analyzes all nucleated cells within a blood sample at single cell resolution. Cells from a patient’s blood sample are deposited on replicate glass slides and compared for morphological features, expression of biomarkers and nuclear integrity, using immunofluorescent staining. Guided by cell phenotype and marker expression, CTCs are individually recovered from the slide surface. Their genomes are amplified (WGA) and analyzed by next generation sequencing (NGS) for the presence of point mutations, copy number alterations, genome wide chromosomal instability, ploidy or genome wide scarring.
- Slide prep: Upon patient blood sample receipt at Epic Sciences, whole blood is lysed and nucleated cells are deposited onto microscope slides.
- Cell staining and CTC identification: Cells are stained for immunofluorescence to identify CTCs and other biomarkers in the study. Coordinates of all CTCs on the slide are preserved.
- Genomics Analysis: Coverslips are removed and cell coordinates are used to relocate CTCs. Once relocated, cells are picked and placed into individual wells of an assay-ready 96 well plate.
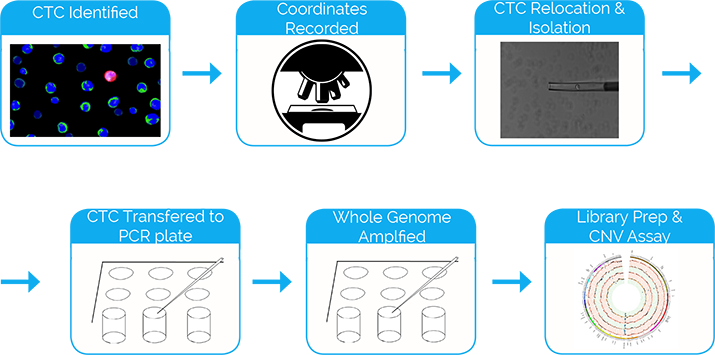
Epic Sciences’ CNV assay service consists of:
- Samples are processed by standard procedures and CTCs are identified (see standard enumeration process for details) for recovery guided by morphology and marker expression.
- Using the unique cell ID number and the associated XY coordinates, CTCs are re-located and recovered from the slide surface.
- Cells are lysed. Their genomes are amplified and constructed into libraries.
- Libraries are sequenced using a HiSeq 2500. Reads are trimmed and aligned.
- Aligned reads are then binned into genomic regions and normalized for variance, read number and WBC control.
- CNV profiles are clustered (k-means and hierarchical) to measure # of clonal populations per patient and associated with cell morphology and biomarker expression to characterize CTC clonality.
Example data:
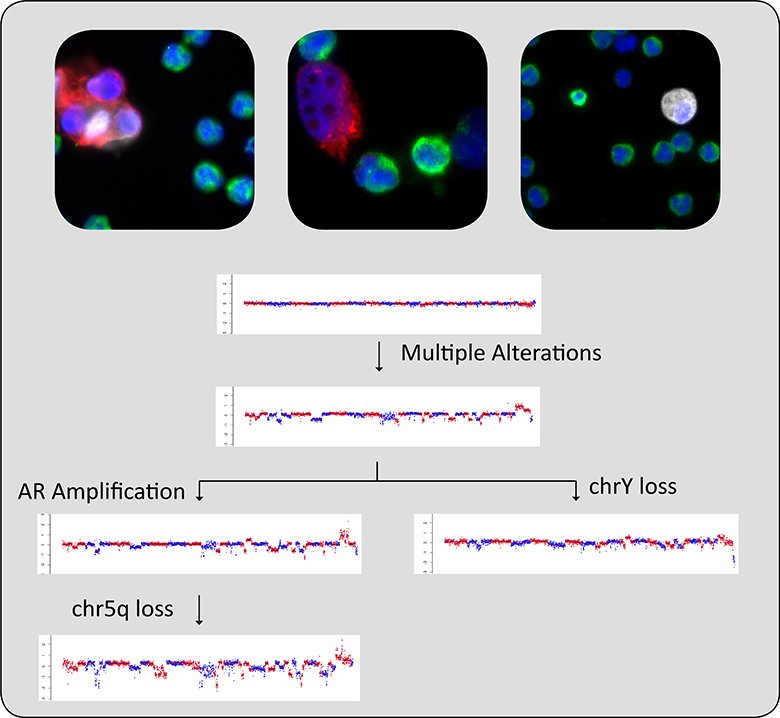
Example data showing the clonal evolution. CTCs from a single patient were identified and characterized through our CNV assay. The distinct clonal species and their relationship to each other are shown in the figure above.
Epic Sciences’ targeted resequencing assay service consists of:
- Samples are processed and CTCs are identified for recovery.
- Using the unique cell ID# and the associated XY coordinates, CTCs are re-located and recovered from slide surface.
- Cells are lysed, their genomes amplified, and constructed into libraries.
- Libraries are target enriched by probe capture for pan-cancer gene exonic regions.
- Following enrichment, libraries are sequenced.
- Reads are aligned and analyzed by single nucleotide variant calling pipeline to characterize the presence on SNV and INDELs in each CTC.
Example data:
Comparison of called variant function (left) and impact (right) across all patient CTCs
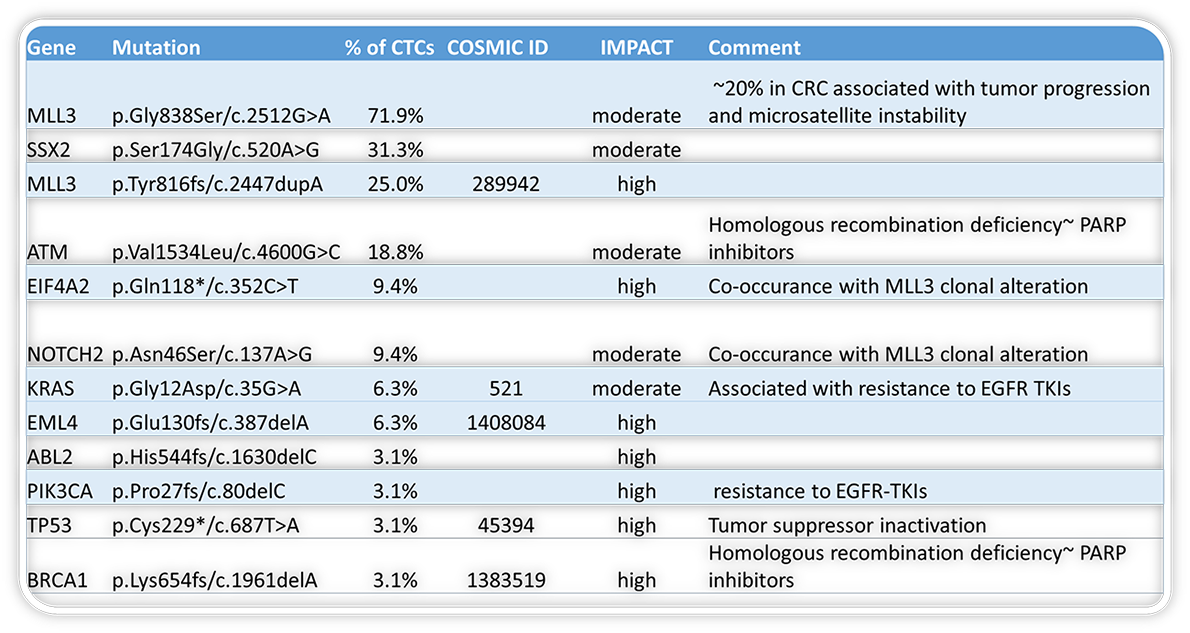
Learn more about our genomics assays are being used:
NAVIGATION
Up to date with Epic Sciences
Learn how these genomic assays are being used to understand AR signaling inhibitor resistance.
